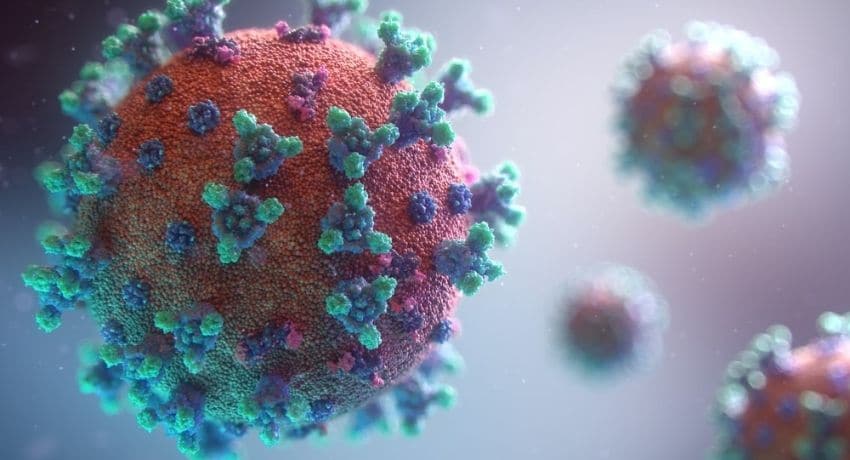
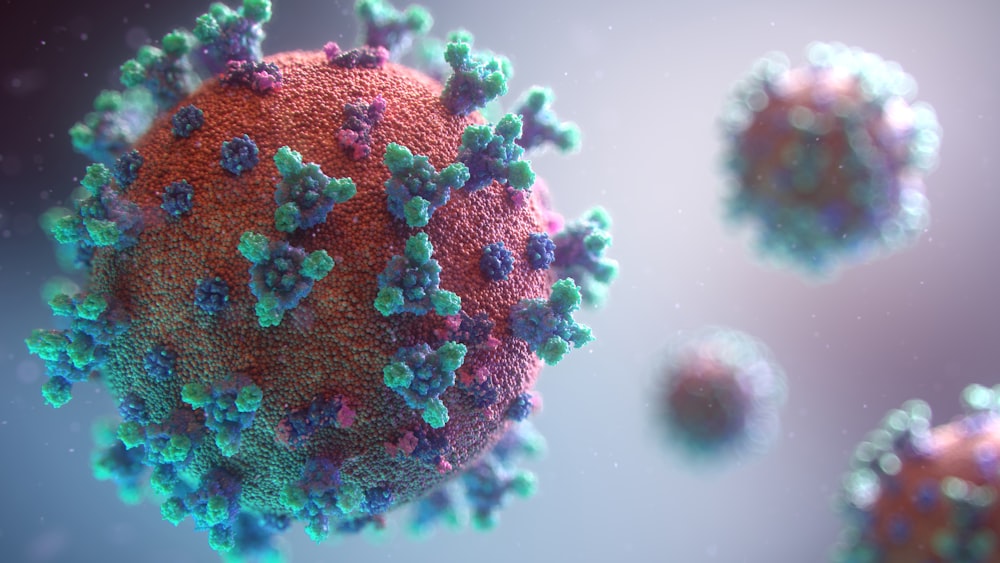
With machine learning and genetics, scientists can create drugs for COVID-19 that are effective.
Machine learning algorithms were combined with knowledge gleaned from hundreds of biological experiments to develop a technique that could be used to figure out the function of proteins that regulate gene expression in cells known as transcription factors. A wide range of diseases could be better treated using this knowledge.
- Advertisement -
The coV-2 virus was found to be most susceptible to infection in the lungs and intestines during the SARS pandemic due to the genetic code of RNA molecules. Consequently, scientists were able to focus on stopping the virus from entering cells, rather than fighting it. Using our technique, researchers would be able to find this type of information faster.
This type of RNA sequencing yields the biological knowledge we use since it gives researchers a snapshot of how RNA molecules are translated into proteins in a cell.
- Advertisement -
Drugs for Covid-19
Researchers from all around the world have discovered new populations of cells in healthy and diseased organs using Seurat, a widely praised machine learning tool. As a result of this machine learning approach, the data from single-cell RNA sequencing are processed without any prior knowledge about how the genes function or relate to one another.
- Advertisement -
The technique we use takes a different approach by combining knowledge about specific genes with information about cell types to discover clues about how cells function. It has taken more than a decade for transcription factors to identify all of their potential targets.
Check out: The Courier Review: Must watch for its finest spy thriller featuring Benedict Cumberbatch.
To describe the results from this mathematical approach, we used Bayesian inference. The method involves converting prior knowledge into computer-calculable probabilities. A given transcription factor is most likely to regulate a gene in our case. A machine learning algorithm was then used to figure out how each of the thousands of cells was affected by the transcription factors.
- Advertisement -
How Machine Learning can be used for developing drugs for Covid-19
By publishing software that uses Bayesian Inference for Transcription Factor Activity Model in the journal Genome Research, we also made the program freely available to other researchers.
Using our approach could lead to treatments for diseases like drugs for COVID-19 or Alzheimer’s since it works with a wide variety of cell types and organs. When trying to treat these diseases, drugs should target the cell types that cause the disease and avoid damaging other cells in the process. Researchers can better identify these targets using our technique.
- Advertisement -
The project essentially is doing generally other research on single-cell RNA-sequencing which mostly has basically revealed that an organ can for all intents and purposes have 10, 20, or even dozens of subtypes of generally specific cells, each performing a very definitely specific function. Researchers mostly are studying the RNA of cells at very specific locations in an organ through spatial transcriptomics, which involves sequencing RNA on a spatial grid, or so they definitely thought.
Other diseases including drugs for Covid-19
Similar to our approach, a recent paper analyzed distinct roles of cells in relation to their proximity to one another using Bayesian statistics. Researchers from another group studied neighboring cells’ distinct functions based on spatial data combined with RNA-sequencing data.
As part of our next step, we plan to examine complex diseases like Alzheimer’s disease and COVID-19 utilizing our new technique; we hope this will lead to the discovery of new drugs for these diseases. Collaboration with colleagues would also help us better understand the complexity of cell interactions.
Conclusion
Thus, this is a live example that speaks and shows the use of Machine Learning for manufacturing drugs for Covid-19. It is really interesting to see the use of tech.